Enhanced decolourisation and degradation of azo dyes using wild versus mutagenic improved bacterial strain: a review
Bhayana Tanya, Saxena Ambika, Gupta Sarika, Dubey Ashish Kumar
Review Articles | Published: 19 December, 2022
First Page: 28
Last Page: 37
Views: 3955
Keywords: Azo dyes, Bioremediation, Mutation, Textile effluent
Abstract
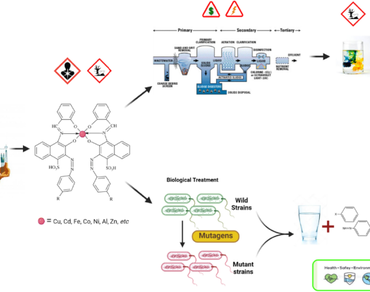
Rapid industrialization over the last few decades has greatly impacted the environment, as various industries release effluent (textile dyeing/printing) which contains copious amount of xenobiotic compound such as azo dyes, salts, organic pollutants, metals, etc. The recalcitrant azo dyes present in textile effluent cause great impact on biota thereby disturbing the integrity of ecosystem. The wastewater discharged into water bodies leads to decline in flora-fauna prevalence due to alteration of the physiochemical characteristics of water (increment in the BOD/COD). Although various physico-chemical methods used for effluent treatment are effective; but are often associated with secondary disposal problem due to the formation of concentrated toxic sludge and presence of toxicants. In recent years, microbial biodegradation has emerged as highly promising approach for textile effluent treatment of xenobiotic toxicants. The indigenous bacteria contain pool of oxidoreductive enzymes (viz. azoreductase, laccase, peroxidase, etc.) that utilize complex xenobiotic compounds of the dyestuffs as substrates and break them into non-toxic by-products. However, limited studies have been reported citing the enhancement in the dye degradation and detoxification efficacy of bacterial species by random mutagenesis approach for the efficient bioremediation of wide spectrum of pigments and dyes. The current review presents the untapped potential of mutagenic approach. It highlights the need by a comprehensive account on the exploration of physical (UV, gamma) and chemical (viz. ethidium bromide, ethyl methane sulfonate) mutagenic agents, respectively. These mutagens alter the genomic profile of the bacteria and facilitate them with higher biodegradation and detoxification efficacies by upregulating the production of oxidoreductive enzymes. The study narrates the prospective for utilization of the wild versus mutant approach consecutively for the mitigation of wide spectrum of azodyes/pigments based on experimental modulations. Review suggests recommendations for the development of novel mutant strains for effective bioremediation strategy as per the permissible compliance limits of pollution board. The approach will thereby able to deal with environmentally noxious pollutants turning one of the most polluted textile industry among the red list to green industry through translational technology.
References
Allam N (2017) Bioremediation efficiency of heavy metals and azo dyes by individual or consortium bacterial species either as free or immobilized cells: a comparative study. Egypt J Bot 57(3):555–564
Arivoli A, Sathiamoorthi T, Satheeshkumar M (2017) Treatment of textile effluent by phyto remediation with aquatic plants: Alternanthera sessilis. Bioremediation and sustainable technologies for cleaner environment. Environ Sci Eng. https://doi.org/10.1007/978-3-319-48439-6_15
Aubert S, Schwitzguebel JP (2004) Screening of plant species for the phytotreatment of wastewater containing sulphonated anthraquinones. Water Res 38:3569
Bechtold T, Turcanu A, Ganglberger E, Geissler S (2003) Natural dyes in modern textile dyehouses—how to combine experiences of two centuries to meet the demands of the future? J Clean Prod. https://doi.org/10.1016/S0959-6526(02)00077-X
Benkhaya S, M’rabet S, Harfi AE (2020) Classifications, properties, recent synthesis and applications of azo dyes. Heliyon 6(1):e03271. https://doi.org/10.1016/j.heliyon.2020.e03271
Brüschweiler BJ, Merlot C (2017) Azo dyes in clothing textiles can be cleaved into a series of mutagenic aromatic amines which are not regulated yet. Regul Toxicol Pharmacol 88:214–226. https://doi.org/10.1016/j.yrtph.2017.06.012
Bu T, Yang R, Zhang Y, Cai Y, Tang Z, Li C, Wu Q, Chen H (2020) Improving decolorization of dyes by laccase from Bacillus licheniformis by random and site-directed mutagenesis. PeerJ 8:e10267. https://doi.org/10.7717/peerj.10267
Chang JS, Kuo TS, Chao YP et al (2000) Azo dye decolorization with a mutant Escherichia coli strain. Biotechnol Lett 22:807–812. https://doi.org/10.1023/A:1005624707777
Chen H (2006) Recent advances in azo dye degrading enzyme research. Curr Protein Pept Sci 7(2):101–111. https://doi.org/10.2174/138920306776359786, https://doi.org/10.1016/j.ecoinf.2015.12.001
Elsahida K, Fauzi A, Saillah I, Siregar I (2019) Sustainability of the use of natural dyes in the textile industry. IOP Conf Ser Earth Environ Sci 399:012065. https://doi.org/10.1088/1755-1315/399/1/012065
Enayatizamir N, Tabandeh F, Rodríguez-Couto S, Yakhchali B, Alikhani HA, Mohammadi L (2011) Biodegradation pathway and detoxification of the diazo dye Reactive Black 5 by Phanerochaete chrysosporium. Bioresour Technol 102(22):10359–10362. https://doi.org/10.1016/j.biortech.2011.08.130 (Epub 2011 Sep 8 PMID: 21955876)
Ergene A, Ada K, Tan S, Katircioglu H (2009) Removal of Remazol Brilliant Blue R dye from aqueous solutions by adsorption onto immobilized Scenedesmus quadricauda: equilibrium and kinetic modeling studies. Desalination 249:1308–1314. https://doi.org/10.1016/j.desal.2009.06.027
Eskandari F, Shahnavaz B, Mashreghi M (2019) Optimization of complete RB-5 azo dye decolorization using novel cold-adapted and mesophilic bacterial consortia. J Environ Manag 241:91–98
Gopinath KP, Murugesan S, Abraham J, Muthukumar K (2009) Bacillus sp. mutant for improved biodegradation of Congo red: random mutagenesis approach. Bioresour Technol 100(24):6295–6300. https://doi.org/10.1016/j.biortech.2009.07.043
Ishchi T, Sibi G (2020) Azo dye degradation by Chlorella vulgaris: optimization and kinetics. Int J Biol Chem 14:1–7
Jamee R, Siddique R (2019) Biodegradation of synthetic dyes of textile effluent by microorganisms: an environmentally and economically sustainable approach. Eur J Microbiol Immunol 9(4):114–118
Jinqi L, Houtian L (1992) Degradation of azo dyes by algae. Environ Pollut (barking, Essex: 1987) 75(3):273–278. https://doi.org/10.1016/0269-7491(92)90127-v
Joshi SM, Inamdar SA, Jadhav JP, Govindwar SP (2013) Random UV mutagenesis approach for enhanced biodegradation of sulfonated azo dye, Green HE4B. Appl Biochem Biotechnol 169(5):1467–1481. https://doi.org/10.1007/s12010-012-0062-5 (Epub 2013 Jan 15, PMID: 23315264)
Kalme S, Ghodake G, Govindwar S (2007) Red HE7B degradation using desulfonation by Pseudomonas desmolyticum NCIM 2112. Int Biodeterior Biodegradation 60:327–333. https://doi.org/10.1016/j.ibiod.2007.05.006
Khehra MS, Saini HS, Sharma DK, Chadha BS, Chimni SS (2006) Biodegradation of azo dye C.I. acid red 88 by an anoxic–aerobic sequential bioreactor. Dyes Pigments 70:1
Křížová H (2015) Natural dyes: their past, present, future and sustainability. In: Recent developments in fibrous material science. O.P.S. Kanina, pp 59–71
Kumar SS, Shantkriti S, Muruganandham T, Murugesh E, Rane N, Govindwar SP (2015) Bioinformatics aided microbial approach for bioremediation of wastewater containing textile dyes. Ecol Inf 31:12–121. https://doi.org/10.1016/j.ecoinf.2015.12.001
Lone TA, Revathi C, Lone RA (2015) Isolation of dye degrading bacillus species from the soil near dyeing industry and its potential application in dye effluent treatment. American-Eurasian J Toxicol Sci 7(3):129–135. https://doi.org/10.5829/idosi.aejts.2015.7.3.94162
Madhavi S, Revankar S, Lela S (2007) Synthetic dye decolorization by white rot fungus, Ganoderma sp. WR1. Bioresour Technol 98:775–780. https://doi.org/10.1016/j.biortech.2006.03.020
Mahmood S, Khalid A, Arshad M, Mahmood T, Crowley DE (2016) Detoxification of azo dyes by bacterial oxidoreductase enzymes. Crib Rev Biotechnol 36(4):639–651. https://doi.org/10.3109/07388551.2015.1004518 (Epub 2015 Feb 10, PMID: 25665634)
Mani A, Hameed SAS (2019) Improved bacterial-fungal consortium as an alternative approach for enhanced decolourisation and degradation of azo dyes: a review. Nat Environ Pollut Technol 18(1):49–64
Maniyam MN, Ibrahim AL, Cass AE (2020) Decolourization and biodegradation of azo dye methyl red by Rhodococcus strain UCC 0016. Environ Technol 41(1):71–85
Moosvi S, Kheer X, Madamwar D (2007) Isolation, characterization and decolorization of textile dyes by a mixed bacterial consortium JW-2. Dyes Pigments 74:723–729. https://doi.org/10.1016/j.dyepig.2006.05.005
Pandey A, Singh P, Iyana L (2007) Bacterial decolorization and degradation of azo dyes. Int Biodeterior Biodegrad 59(2):73–84. https://doi.org/10.1016/j.ibiod.2006.08.006
Pinheiro LRS, Gradíssimo DG, Xavier LP, Santos AV (2022) Degradation of azo dyes: bacterial potential for bioremediation. Sustainability 14(3):1510. https://doi.org/10.3390/su14031510
Pratiwi D, Prasetyo DJ, Dewi C (2019) Adsorption of methylene blue dye using marine algae Ulva lactuca. IOP Conf Ser Earth Environ Sci 251:012012. https://doi.org/10.1088/1755-1315/251/1/012012
Pushpa V, Yogendra K, Mahadevan KM, Mahesh M (2020) Enhancement of azo dye degradation by chemical and physical mutagenesis in identified bacillus: an influential source of industrial dye degradation. Int J Curr Microbiol Appl Sci 9(3):1987–1997. https://doi.org/10.20546/ijcmas.2020.903.231
Ramalho PA, Cardoso MH, Cavaco-Paulo A, Ramalho MT (2004) Characterization of azo reduction activity in a novel ascomycete yeast strain. Appl Environ Microbiol 70(4):2279–2288. https://doi.org/10.1128/AEM.70.4.2279-2288.2004
Rawat D, Mishra V, Sharma RS (2016) Detoxification of azo dyes in the context of environmental processes. Chemosphere 2016(155):591–605. https://doi.org/10.1016/j.chemosphere.2016.04.068
Saratale RG, Saratale GD, Chang JS, Govindwar SP (2011) Bacterial decolorization and degradation of azo dyes: a review. J Taiwan Inst Chem Eng 42(1):138–157. https://doi.org/10.1016/j.jtice.2010.06.006
Sari IP, Simarani K (2019) Decolorization of selected azo dye by Lysinibacillus fusiformis W1B6: biodegradation optimization, isotherm, and kinetic study biosorption mechanism. Adsorpt Sci Technol 37(5–6):492–508. https://doi.org/10.1177/0263617419848897
Sarkar S, Banerjee A, Halder U et al (2017) Degradation of synthetic azo dyes of textile industry: a sustainable approach using microbial enzymes. Water Conserv Sci Eng 2:121–131. https://doi.org/10.1007/s41101-017-0031-5
Saxena A, Gupta S (2020) Bioefficacies of microbes for mitigation of azo dyes in textile industry effluent: a review. BioResources 15(4):9858–9881
Selvaraj V, Karthika ST, Mansiya C, Alagar M (2021) An over review on recently developed techniques, mechanisms and intermediate involved in the advanced azo dye degradation for industrial applications. J Mol Struct 1224:129195. https://doi.org/10.1016/j.molstruc.2020.129195
Shah MP (2018) Azo dye removal technologies. Austin J Biotechnol Bioeng 5(1):1090
Slama HB, ChenariBouket A, Pourhassan Z, Alenezi FN, Silini A, Cherif-Silini H, Oszako T, Luptakova L, Golińska P, Belbahri L (2021) Diversity of synthetic dyes from textile industries, discharge impacts and treatment methods. Appl Sci 11(14):6255. https://doi.org/10.3390/app11146255
Sriram N, Reetha D, Saranraj P (2013) Biological degradation of reactive dyes by using bacteria isolated from dye effluent contaminated soil. Middle East J Sci Res 17(12):1695–1700
Sundarajoo A, Maniyam MN, Azman H, Abdullah H, Yaacob N (2022) Rhodococcus strain UCC 0010 as green biocatalyst for enhanced biodecolourization of Congo red through response surface methodology. Int J Environ Sci Technol 19(4):3305–3322
Tian F, Wang Y, Guo G, Ding K, Yang F, Wang H et al (2021) Enhanced azo dye biodegradation at high salinity by a halophilic bacterial consortium. Bioresour Technol 326:124749
Tripathi A, Srivastava SK (2011) Ecofriendly treatment of azo dyes: biodecolorization using bacterial strains. Int J Biosci Biochem Bioinf 1(1):37–40. https://doi.org/10.7763/IJBBB.2011.V1.7
Ventura-Camargo BD, Marin-Morales MA (2013) Azo dyes: characterization and toxicity—a review. Text Light Ind Sci Technol (TLIST) 2(2):85–103
Watharkar AD, Khandare RV, Kamble AA, Mulla AY, Govindwar SP, Jadhav JP (2013) Phytoremediation potential of Petunia grandiflora Juss., an ornamental plant to degrade a disperse, disulfonated triphenylmethane textile dye Brilliant Blue G. Environ Sci Pollut Res Int 20(2):939–949. https://doi.org/10.1007/s11356-012-0904-2 (Epub 2012 Apr 20, PMID: 22529004)
Author Information
Department of Bioscience and Biotechnology, Banasthali Vidyapith, Tonk, India