Genetic diversity among different landraces of Pearl millet [Cenchrus americanus (L.) Morrone syn. Pennisetum glaucum (L.) R. Br.]
Choudhary Sumitra Kumari, Shekhawat Smita, Harish, Barupal Meena, Kumar Manoj, Shekhawat Narpat S.
Research Articles | Published: 05 July, 2021
First Page: 919
Last Page: 927
Views: 4189
Keywords: Pearl millet, Landrace, Climate change, Genetic diversity, Polymorphism, Principal component analysis
Abstract
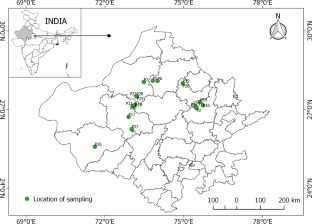
Pearl Millet (PM) is an agronomically important crop, well known for its climate resilient feature and nutritional transcendency. Twenty-eight different tropically adapted Pearl Millet Landraces (PMLRs) of Rajasthan (a north-western state of India) were evaluated for their allelic diversity employing two DNA sequence-based markers: random amplified polymorphic DNA (RAPD) and inter simple sequence repeats (ISSR). After the preliminary screening, 5 RAPD and 5 ISSR primers were identified that gave consistent results across all 28 genotypes. PIC (Polymorphism information content) values were calculated for each primer. Distance matrix revealed that LR (Landrace) P4 shows 100 % dissimilarity with eight LRs (P1, P2, P3, P9, P16, P26, P27 and P28) and may be suitable for breeding programme. Genetic relatedness of these LRs has been estimated using UPGMA (unweighted pair group method with arithmetic mean) and Principal component analysis (PCA) confirm the highly diverse nature of LRs. Results of this study offer possibilities to utilize the genetic diversity for crop improvement and agrobiodiversity conservation.
References
Animasaun DA, Morakinyo JA, Mustapha OT, Krishnamurthy R (2015) Assessment of genetic diversity in accessions of pearl millet (Pennisetum glaucum) and napier grass (Pennisetum purpureum) using microsatellite (ISSR) markers. Iran J Genetics Plant Breeding 4:25–35
Avvall SE (2017) Assessing polymorphism information content (PIC) using SSR molecular markers on local species of Citrullus Colocynthis. Case study: Iran, Sistan-Balouchestan Province. J Mol Biol Res 7:42–49
Basavaraj GL, Murali M, Lavanya SN, Amruthesh KN (2016) Seed priming with biotic agents invokes defense response and enhances plant growth in pearl millet upon infection with Magnaporthe grisea. Biocatal Agric Biotechnol 8:221–227
Bidinger FR, Yadav OP, Rattunde EW (2009) Genetic improvement of pearl millet for the arid zone of northwestern India: lessons from two decades of collaborative ICRISAT-ICAR research. Expl Agric 45:107–115
Busso CS, Devos KM, Ross G, Mortimore M, Adams WM, Ambrose MJ, Alldrick S, Gale MD (2000) Genetic diversity within and among landraces of pearl millet (Pennisetum glaucum) under farmer management in West Africa. Genet Resour Crop Evol 47:561–568
Chesnokov YV, Artemyeva AM (2015) Evaluation of the measure of polymorphism information of genetic diversity. Agric Biol 50:571–578
Choudhary SK, Patel AK, Harish, Shekhawat S, Shekhawat NS (2017) An improved micropropagation system, ex vitro rooting and validation of genetic homogeneity in wild female Momordica dioica: an underutilized nutraceutical vegetable crop. Physiol Mol Biol Plants 23:713–722
Guasmi F, Elfalleh W, Hannachi H, Feres K, Touil L, Marzougui N, Triki T, Ferchichi A (2012) The use of ISSR and RAPD markers for genetic diversity among South Tunisian Barley. Agronomy 952196:1–10
Gupta AK, Harish, Rai MK, Phulwaria M, Shekhawat NS (2011) Isolation of genomic DNA suitable for community analysis from mature trees adapted to arid environment. Gene 487:156–159
Harish, Gupta AK, Phulwaria M, Rai MK, Shekhawat NS (2014) Conservation genetics of endangered medicinal plant Commiphora wightii in Indian Thar Desert. Gene 535(2):266–272
Jaccard P (1908) Nouvelles recherché surla distribution florale. Bull Soc Vaud Sci Nat 44:223–270
Jaiswal S, Antala TJ, Mandavia MK et al (2018) Transcryptomic signature of drought response in pearl millet (Pennisetum glaucum L.) and development of web - genomic resources. Sci Rep 8:1–16
Josiah CC, George DO, Eleazar OM, Nyamu WF (2008) Genetic diversity in Kenyan populations of Acacia senegal (L.) willd revealed by combined RAPD and ISSR markers. Afr J Biotech 7:2333–2340
Jukanti AK, Gaur PM, Gowda CLL, Chibbar RN (2012) Nutritional quality and health benefits of chickpea (Cicer arietinum L.): a review. Br J Nutr 108:11–26
Jukanti AK, Gowda CLL, Rai KN, Manga VK, Bhatt RK (2016) Crops that feed the world. Pearl millet (pennisetumglaucum L.): an important source of food security, nutrition and health in the arid and semi arid tropics. Foods Secur 8:307–329
Kanfany G, Serba DD, Rhode D, Amand PS, Bernardo A, Gangashetty PI, Kane NA, Bai G (2020) Genomic diversity in pearl millet inbred lines derived from landraces and improved varieties. BMC Genom 21:469. https://doi.org/10.1186/s12864-020-06796-4
Karuppanapandian T, Wang HW, Karuppudurai T et al (2010) Efficiency of RAPD and ISSR markers in assessing genetic diversity and relationships in black gram (Vignamungo L. Hepper) varieties. Can J Plant Sci 90:443–452
Khairwal IS, Yadav SK, Rai KN et al (2007) Evaluation and identification of promising pearl millet germplasm for grain and fodder traits. Open Access J Publ ICRISAT 51:1–6
Kumar M, Mishra GP, Singh R, Kumar J, Naik PK, Singh SB (2009) Correspondence of ISSR and RAPD markers for comparative analysis of genetic diversity among different apricot genotypes from cold arid deserts of trans-Himalayas. Physiol Mol Biol Plants 15:225–236
Mohammadi SA, Prasanna BM (2003) Analysis of genetic diversity in crop plants—salient statistical tools and considerations. Crop Sci 43(4):1235–1248
Naino Jika AK, Dussert Y, Raimond C, Garine E, Luxereau A, Takvorian N, Djermakoye RS, Adam T, Robert T (2017) Unexpected pattern of pearl millet genetic diversity among ethno-linguistic groups in the Lake Chad Basin. Heredity 118(5):491–502. doi:https://doi.org/10.1038/hdy.2016.128
Park JR, Yang WT, Kwon YS, Kim HN, Kim KM, Kim DH (2019) Assessment of the genetic diversity of Rice germplasms characterized by black-purple and red pericarp colour using simple sequence repeat markers. Plants 8:1–12
Pavlicek A, Hrda S, Flegr J (1999) Free tree free ware program for construction of phylogenetic trees on the basis of distance data and bootstrap jackknife analysis of the tree Roubstness. Application of the RAPD analysis of genus Frenkelia. FoliaBiol (Praha) 45:97–99
Rai MK, Phulwaria M, Shekhawat NS (2013) Transferability of simple sequence repeat (SSR) markers developed in guava (Psidium guajava L.) to four Myrtaceae species. Mol Biol Rep 40:5067–5071
Reed DH, Frankham R (2001) How closely correlated are molecular and Quantitative measures of genetic variation? A meta-analysis. Evolution 55:1095–1103
Serba DD, Yadav RS (2016) Genomic tools in Pearl Millet breeding for drought tolerance: status and prospects. Front Plant Sci 7:1–10
Sharma YK, Yadav SK, Khairwal IS (2007) Evaluation of pearl millet germplasm lines against downy mildew incited by Sclerospora graminicola in western Rajasthan. SAT ejournal | ejournalicrisatorg 3:1–2
Shekhawat S, Choudhary SK, Gupta AK, Shekhawat NS, Harish, (2013) Determination of genetic diversity of the Morinda tinctoria population in Historical Mandore garden. Proc Natl Acad Sci India Sect B Biol Sci. 83(3):367–370
Shivhare R, Lata C (2017) Exploration of genetic and genomic resources for abiotic and biotic stress tolerance in pearl millet. Front Plant Sci 7:1–17
Stich B, Haussmann BIG, Pasam R, Bhosle S, Hash CT, Melchinger AE, Parzies HK (2010) Patterns of molecular and phenotypic diversity in pearl millet [Pennisetum glaucum (L.) R. Br.] from West and Central Africa and their relation to geographical and environmental parameters. BMC Plant Biol 10:1–10
Thomas M, Dawson JC, Goldringer I, Bonneuil C (2011) Seed exchanges, a key to analysze crop diversity dynamics in Farmer- led on farm conservation. Genet Resour Crop Evol 8:321–338
Ullah A, Ahmad A, Khaliq T, Akhtar J (2017) Recognizing production option for pearl millet in Pakistan under changing climate scenarios. J Integr Agric 16:762–773
URL – 1 Food and Agriculture Organization of the United Nation (2016) FAOSTAT Database. http://www.fao.org/faostat/en/#data/ and accessed on 2 June 2021
Waghmode BT, Sharma KM, Pawar GS, Salunke VD, Kale SP (2018) Genetic diversity and hybrid purity analysis of pearl millet using molecular markers. Int J Curr Microbiol Appl Sci 6:772–779
Yadav OP, Rai KN (2013) Genetic improvement of Pearl millet in India. Agric Res 2:275–292
Author Information
Biotechnology Unit, Department of Botany, UGC, Centre of Advanced Study (CAS), Jai Narain Vyas University, Jodhpur, India